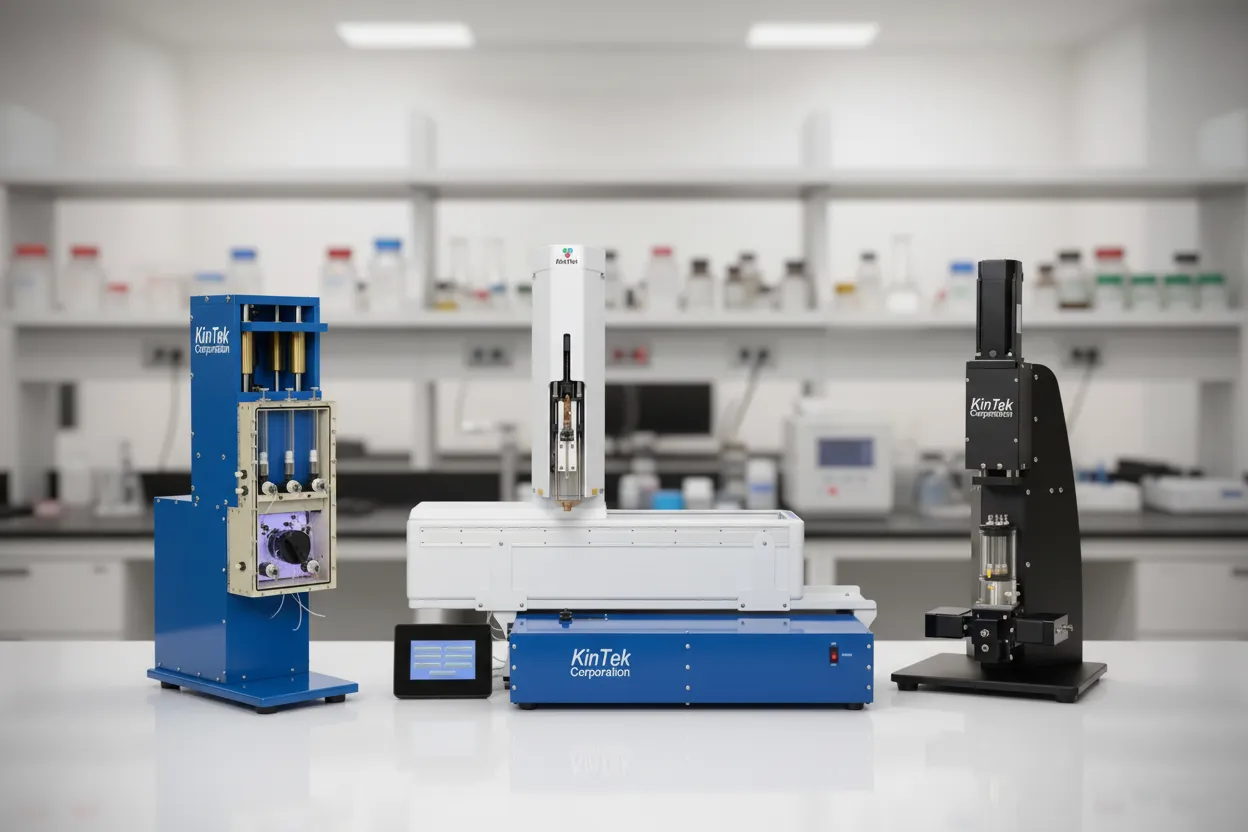
Comprehensive solutions for rapid kinetics analysis
Instruments
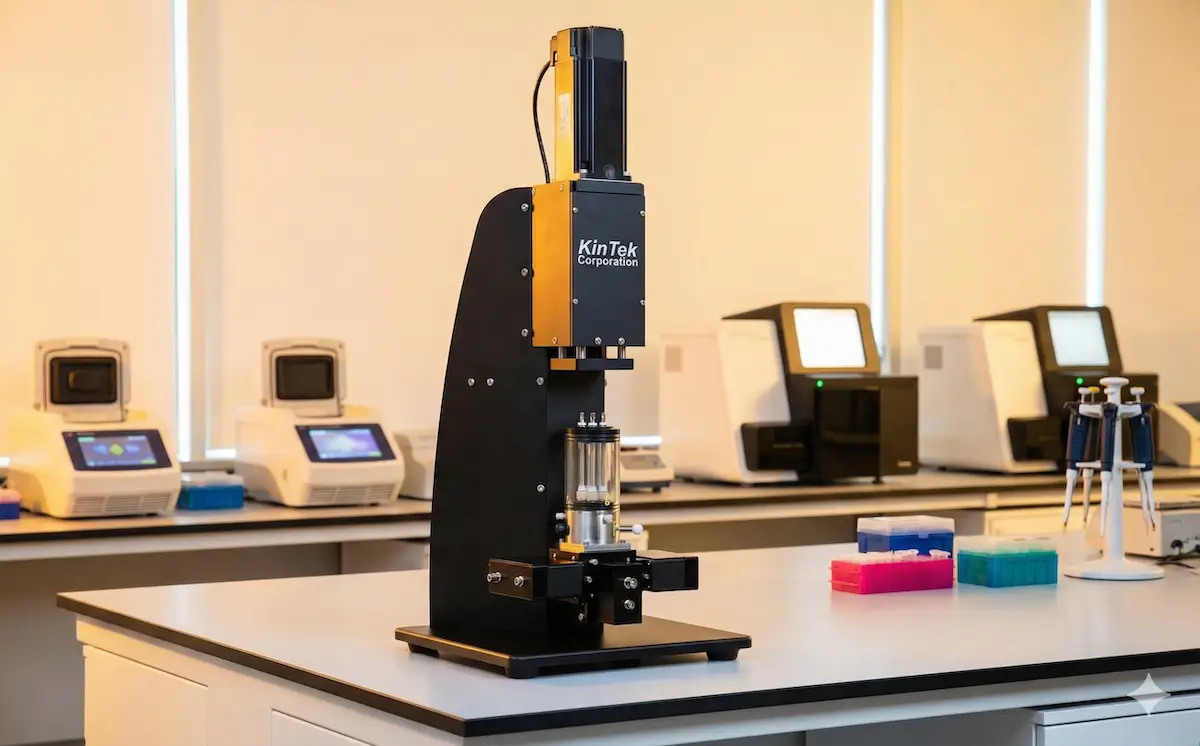
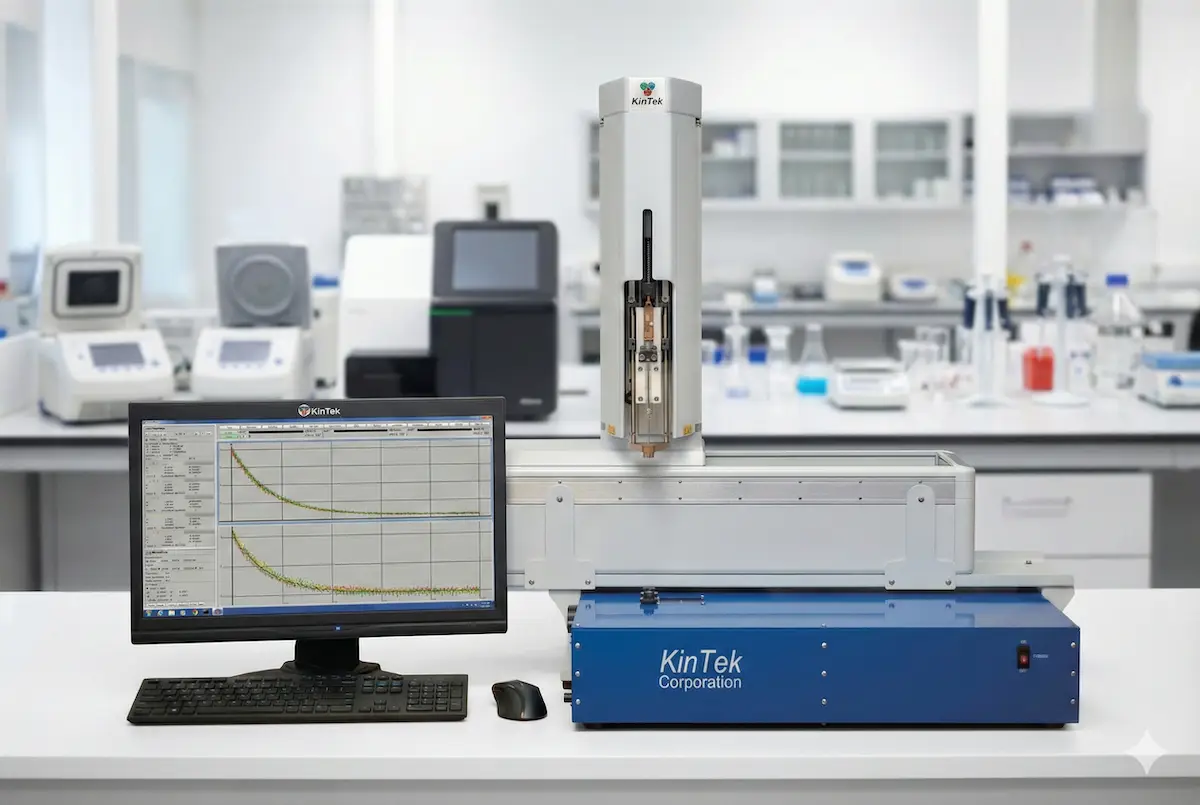
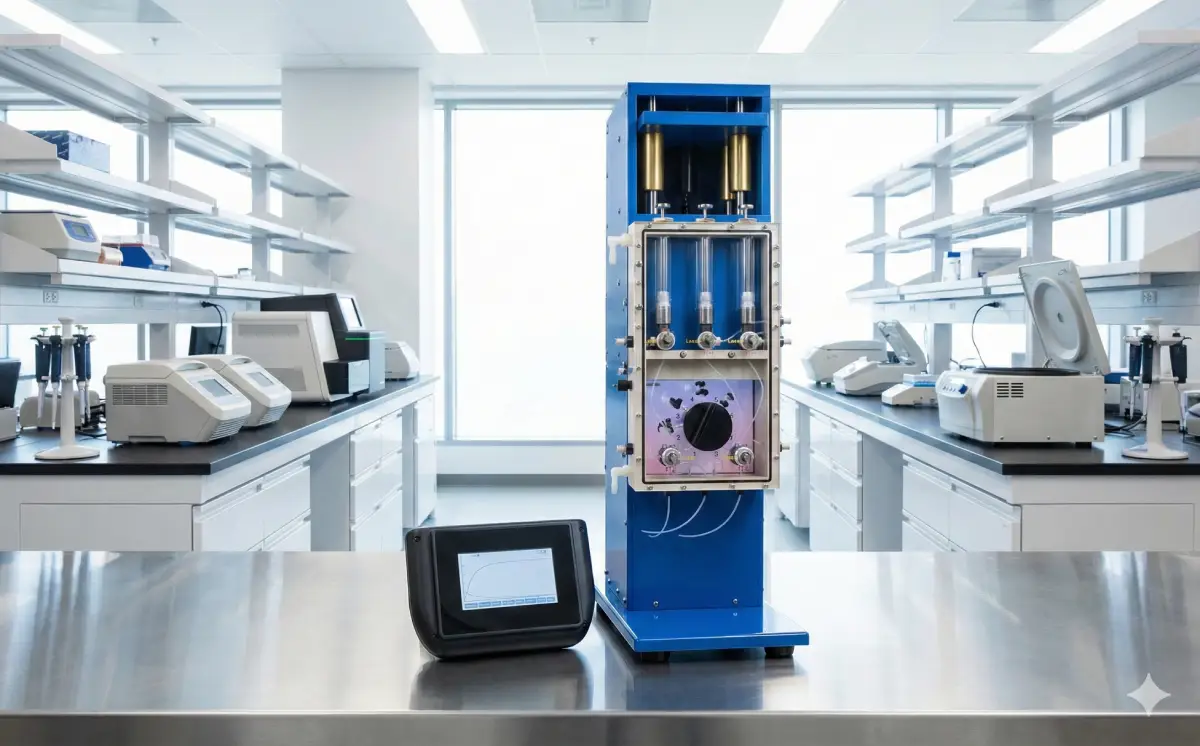
State-of-the-art stopped-flow and quench-flow systems for rapid kinetics measurements.
Software
Advanced kinetic modeling and data analysis software for comprehensive enzyme kinetics studies.
Accessories
Essential components and consumables to optimize your kinetics experiments and workflows.
Books
Educational materials and comprehensive guides for mastering enzyme kinetics principles.
Powering breakthrough research across disciplines
Our rapid kinetics technology enables precise measurements that drive discoveries in enzymology, drug development, and beyond.
Enzyme Kinetics
Uncover reaction mechanisms with millisecond precision
Drug Discovery
Accelerate lead optimization with kinetic insights
Protein Folding
Monitor conformational changes in real-time
Catalysis Research
Characterize catalytic mechanisms with precision
Ready to Advance Your Research?
Join thousands of researchers worldwide who trust KinTek for their kinetics analysis needs.